Introduction & Motivation
A reliable method for early detection for respiratory infections could result in better health outcomes. The gene expression levels of a cell change upon infection and may be a good predictor of illness. These gene expression levels of a cell correspond to the state of the cell and are measured by the abundance of mRNA produced during transcription of each gene. However, for any cell there are tens of thousands of genes measured and limited patient data available. Because of these constraints, we believe a k-nearest classifier would work well to classify infection and we seek to answer the question of which k values and distance measures perform best.
Solution
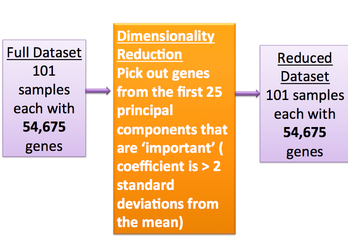
- We first use principal component analysis to reduce the dimensionality of the gene expression profiles.
- We propose a k-nearest neighbor classifier that uses gene expression profiles of individual patient cells to classify them as infected or uninfected
- We iterate with different k values and different distance measures to find method which works best for this data.
TESTING AND TRAINING
Our data set consisted of profiles of 101 sample cells. 79 were from infected cells. 22 were from uninfected cells. Each expression profile is an array of real numbers corresponding to the expression-levels of each gene tested in the cell. We use principal component analysis to reduce the dimensionality of the gene expression profiles.
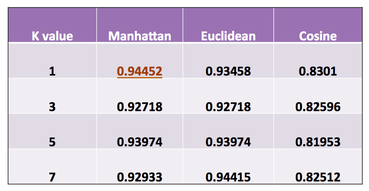
With the reduced data, we train and test a k-nearest classifier using 9-fold validation across k values from 1 to 3 and using L1 and L2 norms as well as cosine similarity as distance measures. We randomized the order of the cells to create an even distribution of infected and uninfected cells per fold.
REsults
Our top performing method was a k-value of 1 and L1 norm at 92.075% accuracy. We evaluated performance using the F1 measure, a weighted ratio of between the rates of precision and recall. The F1 measure was 0.9445. This was an increase of 15.25% in F1 from the lowest performing set (cosine similarity, k = 5). Against a baseline that always chose infected (Fscore = 0.8779) our method provided a 7.5% increase.